Peking University, April 2, 2025: A team from Peking University’s College of Future Technology, led by Professor Xi Peng, in collaboration with Professor Jin Dayong at the Eastern Institute of Technology, has achieved a major breakthrough in imaging 15 cellular structures simultaneously by using lipid membrane probes to label all membrane-bound organelles, combined with dual-color spinning-disk confocal microscopy and deep learning. Their study, titled "Fast Segmentation and Multiplexing Imaging of Organelles in Live Cells", published in Nature Communications (DOI:10.1038/s41467-025-57877-5), utilizes Nile Red dye, spinning-disk confocal microscopy, and deep learning to overcome limitations in traditional fluorescent imaging.
Why it matters:
Cells contain numerous organelles that interact dynamically, but real-time imaging of multiple organelles has been a challenge due to:
• Limited fluorescence labeling: Traditional methods can only label 3-4 organelles due to spectral crosstalk.
• Low efficiency: Increasing the number of labels often fails due to labeling efficiency.
• Short observation windows: Multicolor excitation and multichannel detection cause phototoxicity, limiting long-term imaging.
These limitations have hindered the ability to simultaneously observe organelle interactions, crucial for understanding cellular functions.
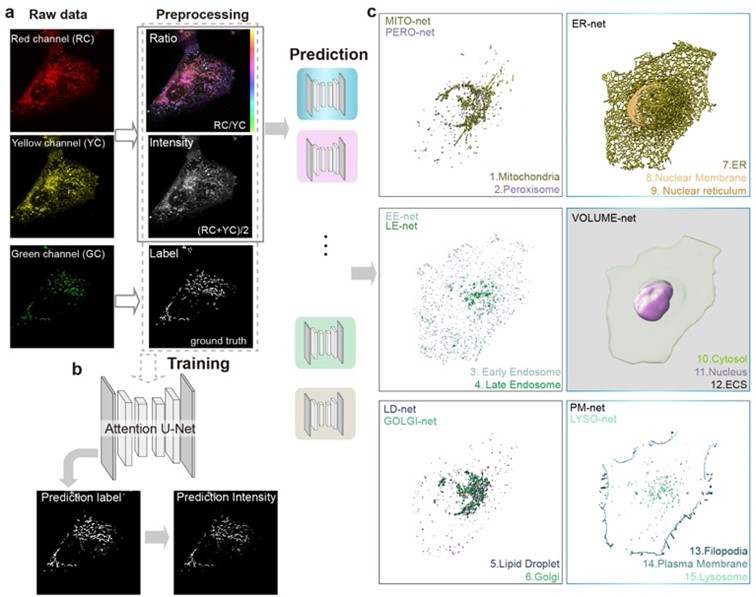 Fig. 1. Precise separation of 15 types of cellular structures within cells through the deep integration of Nile Red lipid dye labeling, super-resolution imaging, and deep learning algorithms: mitochondria, endoplasmic reticulum, lipid droplets, cell membrane, lysosomes, endosomes, Golgi apparatus, nuclear membrane, peroxisomes, cell pseudopodia, nuclear invaginations, cell nucleus, cytoplasm, and extracellular matrix.
Research Methodology
Fig. 1. Precise separation of 15 types of cellular structures within cells through the deep integration of Nile Red lipid dye labeling, super-resolution imaging, and deep learning algorithms: mitochondria, endoplasmic reticulum, lipid droplets, cell membrane, lysosomes, endosomes, Golgi apparatus, nuclear membrane, peroxisomes, cell pseudopodia, nuclear invaginations, cell nucleus, cytoplasm, and extracellular matrix.
Research Methodology
To overcome these challenges, the researchers developed a non-specific labeling, high-throughput imaging strategy:
1. Universal Lipid Staining: Instead of using multiple specific fluorescent labels, they employed Nile Red, a lipid dye that stains all membrane-bound organelles. The dye changes color based on the organelle’s membrane environment, allowing differentiation.
2. Dual-Color Super-Resolution Microscopy: Using spinning-disk confocal microscopy, researchers captured high-speed, low-phototoxicity images of 15 organelles simultaneously (Figure 1).
3. Deep Learning Segmentation: A deep convolutional neural network (DCNN) was trained to recognize and segment organelles automatically based on their optical fingerprints (Figure 2).
This method is:
• Faster and more accurate than traditional segmentation.
• Highly reproducible, allowing large-scale imaging of cellular structures.
This breakthrough technique enables real-time, long-term organelle tracking, significantly improving imaging efficiency while reducing phototoxicity.
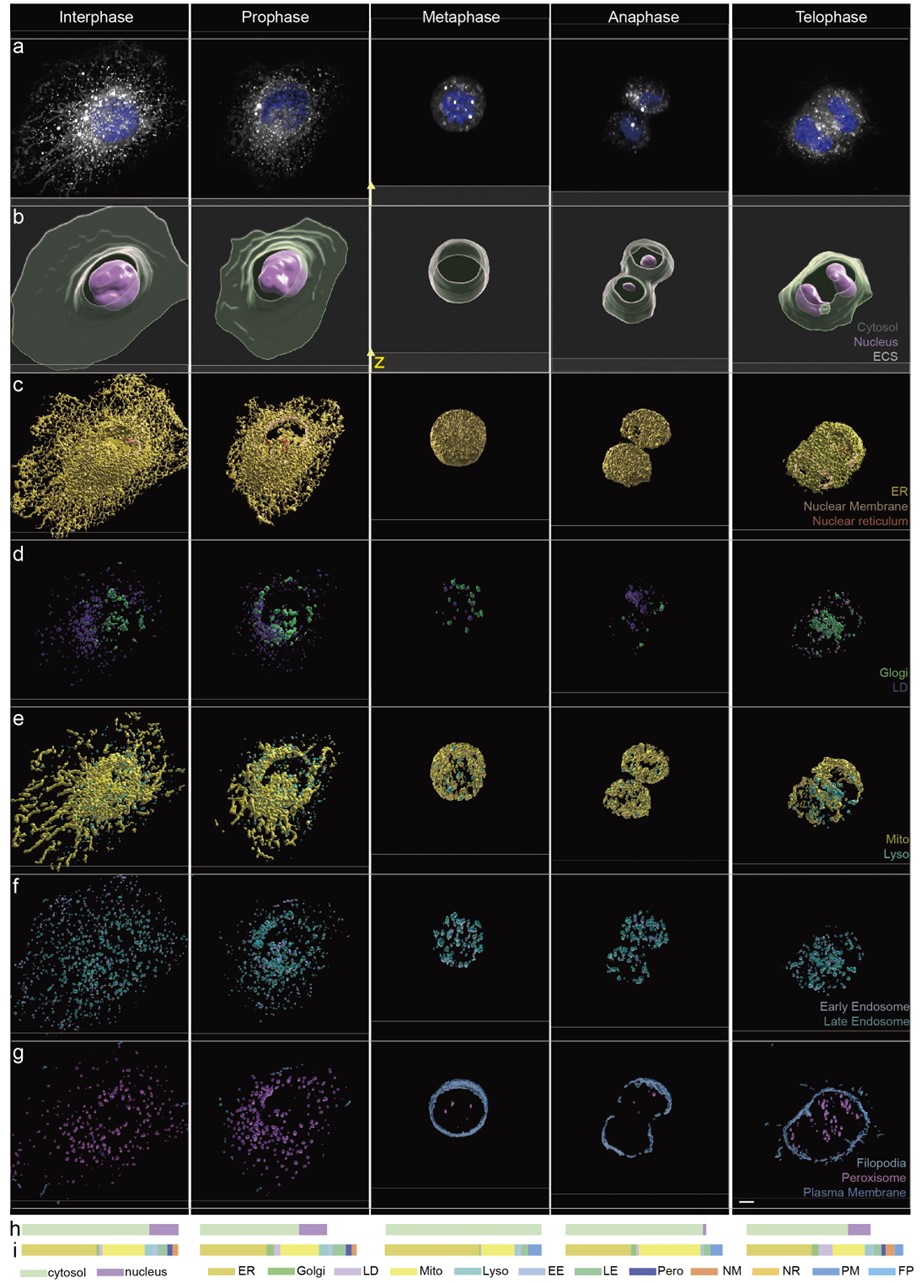 Fig. 2. Panoramic imaging of multiple organelles at different stages of mitosis in cells.
Key Findings
Fig. 2. Panoramic imaging of multiple organelles at different stages of mitosis in cells.
Key Findings
• The technique captured 15 organelles simultaneously such as mitochondria, endoplasmic reticulum, Golgi apparatus, and nucleus (Figure 1).
• Deep learning algorithms were used to automatically segment and predict the location and shape of these organelles, allowing rapid and accurate imaging.
• The method allows long-term observation of organelle dynamics with reduced phototoxicity, making it ideal for studying organellar interactions.
Implications
• This method offers a powerful tool for real-time imaging of organelle interactions in live cells, overcoming the limitations of traditional methods.
• It is also scalable and has been successfully applied to various cell lines and tissues, including fruit fly testis tissue, demonstrating its broad applicability.
*This article is featured in PKU News' "Why It Matters" series. More from this series.
Click “
here” to read the paper.
Written by: Akaash Babar
Edited by: Zhang Jiang
Source: Department of Biomedical Engineering, College of Future Technology, Peking University

